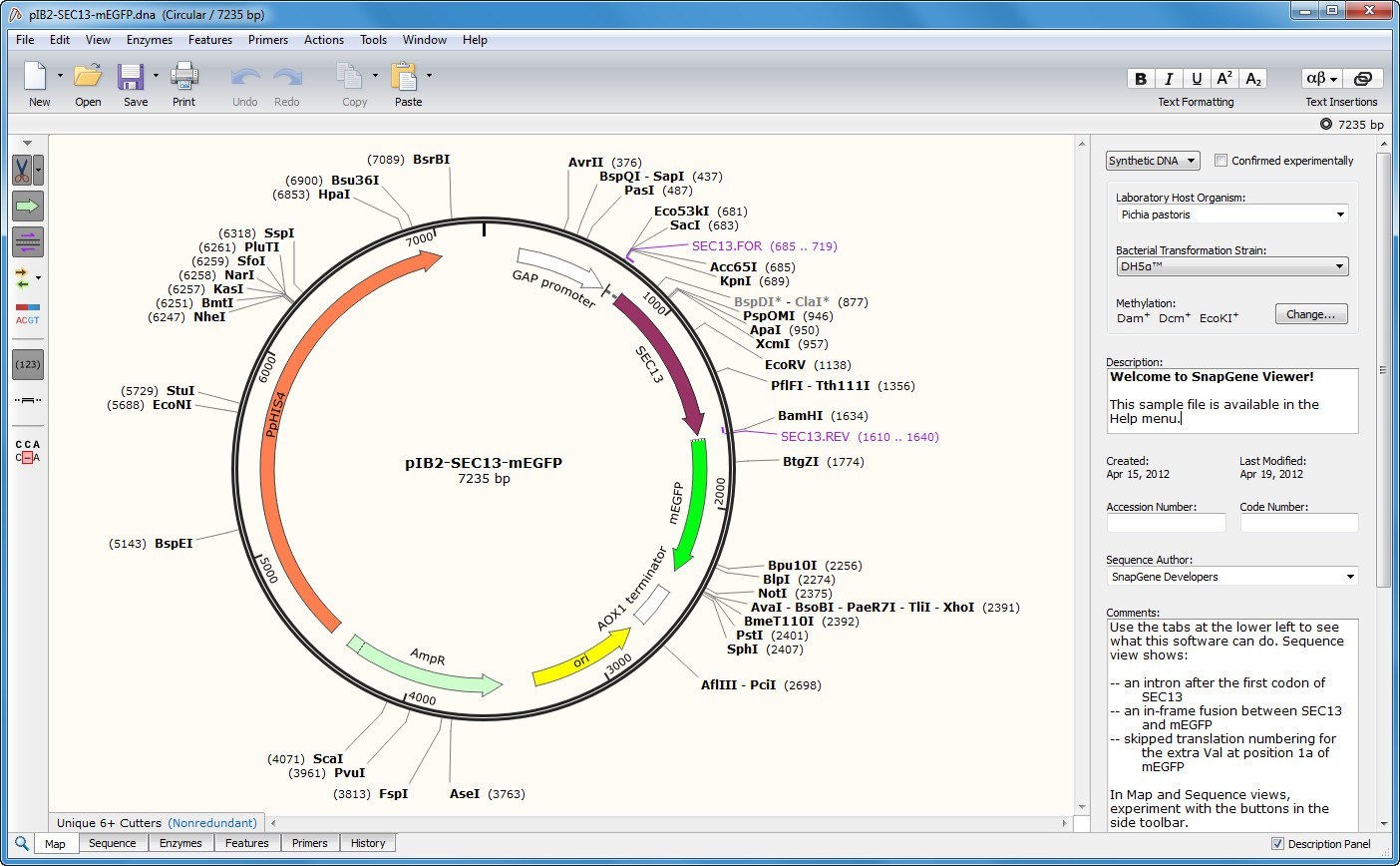
Why two? Because we are cutting a circular, covalently closed piece of DNA. Upon successful cleavage of both strands of DNA using restriction enzymes NdeI and XhoI: We will obtain two, linear DNA fragments. Let’s take a look using this basic diagram or our pET26b plasmid. These are the same bp positions as we determined using the actual sequence. 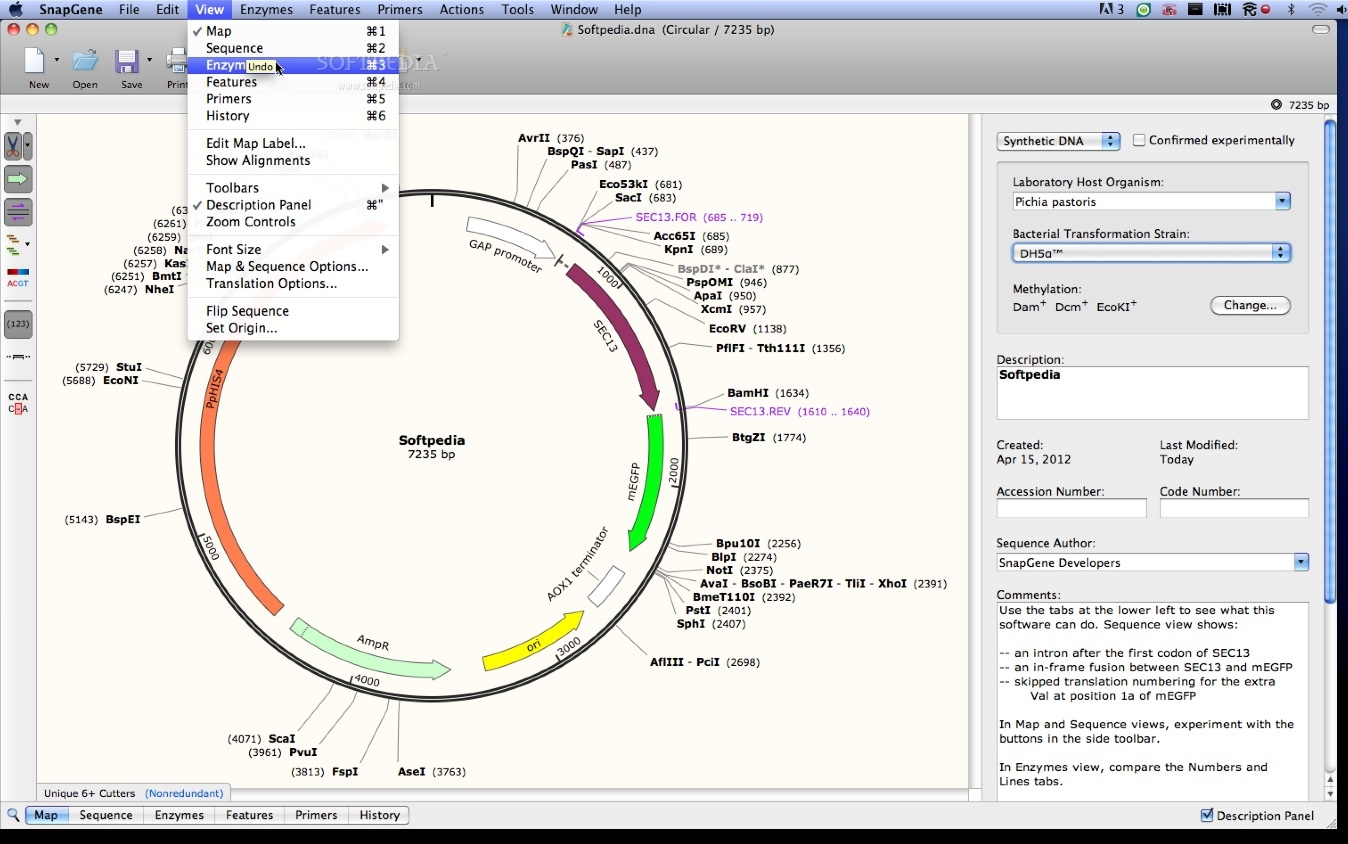
The position of the restriction enzymes (NdeI and XhoI) relative to the T7/lac promoter is clearly indicated… but so are the base pair cut sites! Woohoo!!!!!! Note the numbers in brackets beside each restriction enzyme. Instead, we can take a shortcut based on the information provided in the original, pET26b circular plasmid map shown above. This process for determining DNA fragment sizes works well, but it is time-consuming. We can physically count the base pairs in the yellow highlighted area (there are 128 bps) before we reach the first “c” in the XhoI DNA recognition site, which cuts at exactly bp 5198. If we count base pairs (using the bp numbers on the right as a guide) we can see that the NdeI cuts at exactly the 5070 bp position (remember, we report the bp position relative to the sense strand only). This is to standardize our reporting system, however, we need to keep in mind that we are cleaving both strands of the DNA using restriction enzymes NdeI and XhoI. We typically report the base pair sizes relative to the sense strand (5’ to 3’) only. We can count the number of base pairs (bp) in this area to determine the fragment size cleaved out of the plasmid. The yellow highlighted area contains the DNA fragment cleaved out following NdeI/XhoI digestion of the pET26b plasmid. PET26b plasmid previously digested with NdeI and XhoI =? To answer this question let us zoom in on the pET26b plasmid map and look at the actual sequence surrounding the NdeI and XhoI restriction enzyme DNA recognition sites:
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
